| Synonym | stratifin, HME1, Epithelial cell marker protein 1, SFN, 14-3-3 protein sigma, Stratifin |
|---|
| Refseq ID | NM_006142 |
|---|
| GeneID | 2810 |
|---|
| Unigene ID | Hs.523718 |
|---|
| Probe set ID | 33322_i_at |
|---|
|
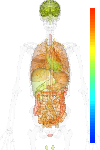 |
| EST |
|
brain:18.60 | |
blood:2.60 | |
connective:115.70 | |
reproductive:9.60 | |
muscular:23.40 | |
alimentary:75.50 | |
liver:- | |
lung:53.80 | |
urinary:- | |
endo/exo-crine:23.30 | |
|
| GeneChip |
|
brain:7.22 | |
blood:8.44 | |
connective:9.40 | |
reproductive:8.48 | |
muscular:6.94 | |
alimentary:10.30 | |
liver:7.06 | |
lung:8.86 | |
urinary:7.23 | |
endo/exo-crine:8.74 | |
|
| CAGE |
|
brain:0.49 | |
blood:2.08 | |
connective:0.36 | |
reproductive:2.17 | |
muscular:0.06 | |
alimentary:4.46 | |
liver:0.55 | |
lung:2.90 | |
urinary:2.85 | |
endo/exo-crine:0.89 | |
|
| RNA-seq |
|
brain:1.29 | |
blood:0.31 | |
connective:1.50 | |
reproductive:1.43 | |
muscular:0.03 | |
alimentary:2.64 | |
liver:0.01 | |
lung:5.48 | |
urinary:1.34 | |
endo/exo-crine:1.40 | |
|
| Synonym | cell death-inducing DFFA-like effector b, CIDEB, Cell death-inducing DFFA-like effector B, Cell death activator CIDE-B |
|---|
| Refseq ID | NM_014430 |
|---|
| GeneID | 27141 |
|---|
| Unigene ID | - |
|---|
| Probe set ID | 221188_s_at |
|---|
|
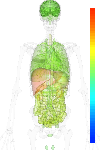 |
| EST |
nodata |
|
| GeneChip |
|
brain:6.16 | |
blood:6.74 | |
connective:6.72 | |
reproductive:6.20 | |
muscular:6.44 | |
alimentary:6.90 | |
liver:9.92 | |
lung:6.51 | |
urinary:7.89 | |
endo/exo-crine:6.71 | |
|
| CAGE |
|
brain:1.17 | |
blood:2.55 | |
connective:2.28 | |
reproductive:1.94 | |
muscular:1.38 | |
alimentary:4.18 | |
liver:6.64 | |
lung:2.03 | |
urinary:3.02 | |
endo/exo-crine:1.86 | |
|
| RNA-seq |
|
brain:2.52 | |
blood:3.29 | |
connective:2.87 | |
reproductive:3.26 | |
muscular:2.60 | |
alimentary:2.71 | |
liver:6.76 | |
lung:2.64 | |
urinary:4.45 | |
endo/exo-crine:3.10 | |
|
| Synonym | cell death-inducing DFFA-like effector b, CIDEB, Cell death-inducing DFFA-like effector B, Cell death activator CIDE-B |
|---|
| Refseq ID | NM_014430 |
|---|
| GeneID | 27141 |
|---|
| Unigene ID | Hs.642693 |
|---|
| Probe set ID | 221188_s_at |
|---|
|
 |
| EST |
|
brain:22.80 | |
blood:36.50 | |
connective:35.30 | |
reproductive:23.30 | |
muscular:- | |
alimentary:30.20 | |
liver:- | |
lung:34.20 | |
urinary:9.60 | |
endo/exo-crine:3.90 | |
|
| GeneChip |
|
brain:6.16 | |
blood:6.74 | |
connective:6.72 | |
reproductive:6.20 | |
muscular:6.44 | |
alimentary:6.90 | |
liver:9.92 | |
lung:6.51 | |
urinary:7.89 | |
endo/exo-crine:6.71 | |
|
| CAGE |
|
brain:1.17 | |
blood:2.55 | |
connective:2.28 | |
reproductive:1.94 | |
muscular:1.38 | |
alimentary:4.18 | |
liver:6.64 | |
lung:2.03 | |
urinary:3.02 | |
endo/exo-crine:1.86 | |
|
| RNA-seq |
|
brain:2.52 | |
blood:3.29 | |
connective:2.87 | |
reproductive:3.26 | |
muscular:2.60 | |
alimentary:2.71 | |
liver:6.76 | |
lung:2.64 | |
urinary:4.45 | |
endo/exo-crine:3.10 | |
|
| Synonym | stratifin, HME1, Epithelial cell marker protein 1, SFN, 14-3-3 protein sigma, Stratifin |
|---|
| Refseq ID | NM_006142 |
|---|
| GeneID | 2810 |
|---|
| Unigene ID | Hs.523718 |
|---|
| Probe set ID | 33323_r_at |
|---|
|
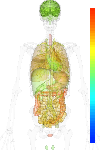 |
| EST |
|
brain:18.60 | |
blood:2.60 | |
connective:115.70 | |
reproductive:9.60 | |
muscular:23.40 | |
alimentary:75.50 | |
liver:- | |
lung:53.80 | |
urinary:- | |
endo/exo-crine:23.30 | |
|
| GeneChip |
|
brain:6.56 | |
blood:7.54 | |
connective:8.54 | |
reproductive:7.64 | |
muscular:6.53 | |
alimentary:9.68 | |
liver:6.54 | |
lung:8.10 | |
urinary:6.60 | |
endo/exo-crine:7.95 | |
|
| CAGE |
|
brain:0.49 | |
blood:2.08 | |
connective:0.36 | |
reproductive:2.17 | |
muscular:0.06 | |
alimentary:4.46 | |
liver:0.55 | |
lung:2.90 | |
urinary:2.85 | |
endo/exo-crine:0.89 | |
|
| RNA-seq |
|
brain:1.29 | |
blood:0.31 | |
connective:1.50 | |
reproductive:1.43 | |
muscular:0.03 | |
alimentary:2.64 | |
liver:0.01 | |
lung:5.48 | |
urinary:1.34 | |
endo/exo-crine:1.40 | |
|
| Synonym | CRIP, hCRHP, cysteine-rich protein 1 (intestinal), Cysteine-rich protein 1, Cysteine-rich heart protein, CRIP1, CRHP, Cysteine-rich intestinal protein |
|---|
| Refseq ID | NM_001311 |
|---|
| GeneID | 1396 |
|---|
| Unigene ID | Hs.70327 |
|---|
| Probe set ID | 205081_at |
|---|
|
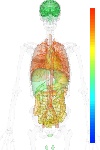 |
| EST |
|
brain:2.50 | |
blood:13.00 | |
connective:6.40 | |
reproductive:15.00 | |
muscular:15.60 | |
alimentary:20.10 | |
liver:- | |
lung:14.70 | |
urinary:9.60 | |
endo/exo-crine:3.90 | |
|
| GeneChip |
|
brain:5.22 | |
blood:8.50 | |
connective:8.92 | |
reproductive:6.82 | |
muscular:6.89 | |
alimentary:8.20 | |
liver:5.11 | |
lung:9.09 | |
urinary:6.01 | |
endo/exo-crine:7.32 | |
|
| CAGE |
|
brain:1.24 | |
blood:4.30 | |
connective:3.62 | |
reproductive:3.04 | |
muscular:3.99 | |
alimentary:4.68 | |
liver:1.07 | |
lung:4.43 | |
urinary:3.16 | |
endo/exo-crine:2.34 | |
|
| RNA-seq |
|
brain:2.09 | |
blood:6.96 | |
connective:5.38 | |
reproductive:5.35 | |
muscular:4.19 | |
alimentary:5.69 | |
liver:1.67 | |
lung:8.17 | |
urinary:4.74 | |
endo/exo-crine:5.76 | |
|
| Synonym | MGC23427, BTBD14A, BTB (POZ) domain containing 14A, BTBD14 |
|---|
| Refseq ID | NM_144653 |
|---|
| GeneID | 138151 |
|---|
| Unigene ID | Hs.112895 |
|---|
| Probe set ID | 212993_at |
|---|
|
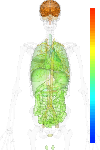 |
| EST |
|
brain:4.20 | |
blood:- | |
connective:3.20 | |
reproductive:15.00 | |
muscular:- | |
alimentary:25.20 | |
liver:- | |
lung:14.70 | |
urinary:19.20 | |
endo/exo-crine:3.90 | |
|
| GeneChip |
|
brain:8.70 | |
blood:6.12 | |
connective:6.36 | |
reproductive:6.60 | |
muscular:6.43 | |
alimentary:6.25 | |
liver:6.25 | |
lung:6.18 | |
urinary:6.33 | |
endo/exo-crine:6.08 | |
|
| CAGE |
|
brain:2.83 | |
blood:2.50 | |
connective:3.21 | |
reproductive:2.95 | |
muscular:3.04 | |
alimentary:3.34 | |
liver:3.38 | |
lung:3.17 | |
urinary:3.46 | |
endo/exo-crine:2.53 | |
|
| RNA-seq |
|
brain:4.33 | |
blood:2.48 | |
connective:3.12 | |
reproductive:3.56 | |
muscular:2.98 | |
alimentary:3.36 | |
liver:2.75 | |
lung:2.62 | |
urinary:2.58 | |
endo/exo-crine:3.07 | |
|
| Synonym | Mismatch-specific DNA N-glycosylase, MED1, Methyl-CpG-binding domain protein 4, MBD4, methyl-CpG binding domain protein 4, Methyl-CpG-binding endonuclease 1, Methyl-CpG-binding protein MBD4 |
|---|
| Refseq ID | NM_003925 |
|---|
| GeneID | 8930 |
|---|
| Unigene ID | Hs.35947 |
|---|
| Probe set ID | 209579_s_at |
|---|
|
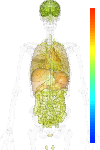 |
| EST |
|
brain:81.90 | |
blood:159.10 | |
connective:93.20 | |
reproductive:101.20 | |
muscular:78.10 | |
alimentary:120.70 | |
liver:52.80 | |
lung:171.10 | |
urinary:105.60 | |
endo/exo-crine:85.40 | |
|
| GeneChip |
|
brain:7.04 | |
blood:8.04 | |
connective:7.25 | |
reproductive:7.69 | |
muscular:7.21 | |
alimentary:7.66 | |
liver:8.28 | |
lung:7.93 | |
urinary:7.54 | |
endo/exo-crine:7.56 | |
|
| CAGE |
|
brain:3.25 | |
blood:3.74 | |
connective:3.46 | |
reproductive:3.19 | |
muscular:3.46 | |
alimentary:3.27 | |
liver:3.71 | |
lung:3.55 | |
urinary:3.51 | |
endo/exo-crine:3.31 | |
|
| RNA-seq |
|
brain:3.28 | |
blood:4.03 | |
connective:3.33 | |
reproductive:3.96 | |
muscular:2.90 | |
alimentary:3.17 | |
liver:3.54 | |
lung:3.10 | |
urinary:4.15 | |
endo/exo-crine:3.80 | |
|
| Synonym | HNPCC, hMLH1 mismatch repair, MGC5172, COCA2, MLH1, HNPCC2, DNA mismatch repair protein Mlh1, FCC2, MutL protein homolog 1, hMLH1 |
|---|
| Refseq ID | NM_000249 |
|---|
| GeneID | 4292 |
|---|
| Unigene ID | Hs.195364 |
|---|
| Probe set ID | 202520_s_at |
|---|
|
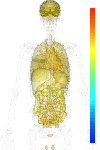 |
| EST |
|
brain:87.80 | |
blood:112.10 | |
connective:125.30 | |
reproductive:116.30 | |
muscular:101.50 | |
alimentary:115.70 | |
liver:84.40 | |
lung:58.70 | |
urinary:67.20 | |
endo/exo-crine:174.70 | |
|
| GeneChip |
|
brain:7.66 | |
blood:7.84 | |
connective:7.75 | |
reproductive:8.16 | |
muscular:8.26 | |
alimentary:7.67 | |
liver:7.54 | |
lung:7.58 | |
urinary:7.51 | |
endo/exo-crine:7.83 | |
|
| CAGE |
|
brain:3.52 | |
blood:3.49 | |
connective:3.35 | |
reproductive:3.48 | |
muscular:3.59 | |
alimentary:3.00 | |
liver:3.35 | |
lung:3.37 | |
urinary:3.24 | |
endo/exo-crine:3.44 | |
|
| RNA-seq |
|
brain:1.79 | |
blood:0.56 | |
connective:2.25 | |
reproductive:2.69 | |
muscular:2.08 | |
alimentary:2.15 | |
liver:1.22 | |
lung:1.18 | |
urinary:1.96 | |
endo/exo-crine:2.91 | |
|
| Synonym | HNPCC, hMLH1 mismatch repair, MGC5172, COCA2, MLH1, HNPCC2, DNA mismatch repair protein Mlh1, FCC2, MutL protein homolog 1, hMLH1 |
|---|
| Refseq ID | NM_001167617 |
|---|
| GeneID | 4292 |
|---|
| Unigene ID | Hs.195364 |
|---|
| Probe set ID | 202520_s_at |
|---|
|
 |
| EST |
|
brain:87.80 | |
blood:112.10 | |
connective:125.30 | |
reproductive:116.30 | |
muscular:101.50 | |
alimentary:115.70 | |
liver:84.40 | |
lung:58.70 | |
urinary:67.20 | |
endo/exo-crine:174.70 | |
|
| GeneChip |
|
brain:7.66 | |
blood:7.84 | |
connective:7.75 | |
reproductive:8.16 | |
muscular:8.26 | |
alimentary:7.67 | |
liver:7.54 | |
lung:7.58 | |
urinary:7.51 | |
endo/exo-crine:7.83 | |
|
| CAGE |
|
brain:3.52 | |
blood:3.49 | |
connective:3.35 | |
reproductive:3.48 | |
muscular:3.59 | |
alimentary:3.00 | |
liver:3.35 | |
lung:3.37 | |
urinary:3.24 | |
endo/exo-crine:3.44 | |
|
| RNA-seq |
|
brain:0.00 | |
blood:2.60 | |
connective:0.00 | |
reproductive:0.75 | |
muscular:0.38 | |
alimentary:0.00 | |
liver:1.08 | |
lung:1.37 | |
urinary:1.31 | |
endo/exo-crine:0.52 | |
|
| Synonym | HNPCC, hMLH1 mismatch repair, MGC5172, COCA2, MLH1, HNPCC2, DNA mismatch repair protein Mlh1, FCC2, MutL protein homolog 1, hMLH1 |
|---|
| Refseq ID | NM_001167618 |
|---|
| GeneID | 4292 |
|---|
| Unigene ID | Hs.195364 |
|---|
| Probe set ID | 202520_s_at |
|---|
|
 |
| EST |
|
brain:87.80 | |
blood:112.10 | |
connective:125.30 | |
reproductive:116.30 | |
muscular:101.50 | |
alimentary:115.70 | |
liver:84.40 | |
lung:58.70 | |
urinary:67.20 | |
endo/exo-crine:174.70 | |
|
| GeneChip |
|
brain:7.66 | |
blood:7.84 | |
connective:7.75 | |
reproductive:8.16 | |
muscular:8.26 | |
alimentary:7.67 | |
liver:7.54 | |
lung:7.58 | |
urinary:7.51 | |
endo/exo-crine:7.83 | |
|
| CAGE |
|
brain:3.52 | |
blood:3.49 | |
connective:3.35 | |
reproductive:3.48 | |
muscular:3.59 | |
alimentary:3.00 | |
liver:3.35 | |
lung:3.37 | |
urinary:3.24 | |
endo/exo-crine:3.44 | |
|
| RNA-seq |
|
brain:2.46 | |
blood:1.03 | |
connective:1.90 | |
reproductive:3.17 | |
muscular:3.24 | |
alimentary:2.32 | |
liver:2.59 | |
lung:1.20 | |
urinary:2.72 | |
endo/exo-crine:2.36 | |
|