| Synonym | Protein MIG-5, Metalloproteinase inhibitor 3 precursor, TIMP3, TIMP-3, K222TA2, Tissue inhibitor of metalloproteinases 3, SFD, HSMRK222, K222 |
|---|
| Refseq ID | NM_000362 |
|---|
| GeneID | 7078 |
|---|
| Unigene ID | Hs.644633 |
|---|
| Probe set ID | 201150_s_at |
|---|
|
 |
| EST |
nodata |
|
| GeneChip |
|
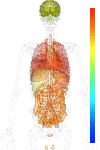
brain:7.37 | |
blood:8.75 | |
connective:10.20 | |
reproductive:9.29 | |
muscular:9.11 | |
alimentary:8.95 | |
liver:7.30 | |
lung:10.79 | |
urinary:9.23 | |
endo/exo-crine:9.36 | |
|
| CAGE |
|
brain:4.64 | |
blood:5.34 | |
connective:7.39 | |
reproductive:6.48 | |
muscular:6.36 | |
alimentary:5.50 | |
liver:4.65 | |
lung:7.24 | |
urinary:6.23 | |
endo/exo-crine:5.49 | |
|
| RNA-seq |
|
brain:5.40 | |
blood:7.39 | |
connective:9.38 | |
reproductive:6.87 | |
muscular:6.46 | |
alimentary:7.88 | |
liver:5.12 | |
lung:9.54 | |
urinary:7.95 | |
endo/exo-crine:8.04 | |
|
| Synonym | Protein MIG-5, Metalloproteinase inhibitor 3 precursor, TIMP3, TIMP-3, K222TA2, Tissue inhibitor of metalloproteinases 3, SFD, HSMRK222, K222 |
|---|
| Refseq ID | NM_000362 |
|---|
| GeneID | 7078 |
|---|
| Unigene ID | - |
|---|
| Probe set ID | 201150_s_at |
|---|
|
 |
| EST |
nodata |
|
| GeneChip |
|
brain:7.37 | |
blood:8.75 | |
connective:10.20 | |
reproductive:9.29 | |
muscular:9.11 | |
alimentary:8.95 | |
liver:7.30 | |
lung:10.79 | |
urinary:9.23 | |
endo/exo-crine:9.36 | |
|
| CAGE |
|
brain:4.64 | |
blood:5.34 | |
connective:7.39 | |
reproductive:6.48 | |
muscular:6.36 | |
alimentary:5.50 | |
liver:4.65 | |
lung:7.24 | |
urinary:6.23 | |
endo/exo-crine:5.49 | |
|
| RNA-seq |
|
brain:5.40 | |
blood:7.39 | |
connective:9.38 | |
reproductive:6.87 | |
muscular:6.46 | |
alimentary:7.88 | |
liver:5.12 | |
lung:9.54 | |
urinary:7.95 | |
endo/exo-crine:8.04 | |
|
| Synonym | RNA-dependent helicase p72, DEAD box protein p72, Probable ATP-dependent RNA helicase DDX17, DEAD box protein 17, DEAD (Asp-Glu-Ala-Asp) box polypeptide 17, RH70, DDX17, DKFZp761H2016, P72 |
|---|
| Refseq ID | NM_001098504 |
|---|
| GeneID | 10521 |
|---|
| Unigene ID | Hs.528305 |
|---|
| Probe set ID | 208718_at |
|---|
|
 |
| EST |
|
brain:366.40 | |
blood:299.90 | |
connective:240.90 | |
reproductive:210.70 | |
muscular:226.40 | |
alimentary:296.80 | |
liver:158.30 | |
lung:268.80 | |
urinary:278.50 | |
endo/exo-crine:252.40 | |
|
| GeneChip |
|
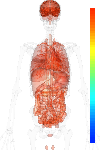
brain:10.40 | |
blood:10.71 | |
connective:10.44 | |
reproductive:10.75 | |
muscular:10.08 | |
alimentary:10.28 | |
liver:10.08 | |
lung:10.77 | |
urinary:10.74 | |
endo/exo-crine:10.60 | |
|
| CAGE |
|
brain:6.00 | |
blood:5.88 | |
connective:5.81 | |
reproductive:5.86 | |
muscular:5.54 | |
alimentary:5.74 | |
liver:4.86 | |
lung:5.94 | |
urinary:5.61 | |
endo/exo-crine:6.05 | |
|
| RNA-seq |
|
brain:3.73 | |
blood:5.90 | |
connective:3.88 | |
reproductive:5.69 | |
muscular:2.71 | |
alimentary:4.88 | |
liver:3.23 | |
lung:2.25 | |
urinary:4.78 | |
endo/exo-crine:4.54 | |
|
| Synonym | RNA-dependent helicase p72, DEAD box protein p72, Probable ATP-dependent RNA helicase DDX17, DEAD box protein 17, DEAD (Asp-Glu-Ala-Asp) box polypeptide 17, RH70, DDX17, DKFZp761H2016, P72 |
|---|
| Refseq ID | NM_006386 |
|---|
| GeneID | 10521 |
|---|
| Unigene ID | Hs.528305 |
|---|
| Probe set ID | 208718_at |
|---|
|
 |
| EST |
|
brain:366.40 | |
blood:299.90 | |
connective:240.90 | |
reproductive:210.70 | |
muscular:226.40 | |
alimentary:296.80 | |
liver:158.30 | |
lung:268.80 | |
urinary:278.50 | |
endo/exo-crine:252.40 | |
|
| GeneChip |
|
brain:10.40 | |
blood:10.71 | |
connective:10.44 | |
reproductive:10.75 | |
muscular:10.08 | |
alimentary:10.28 | |
liver:10.08 | |
lung:10.77 | |
urinary:10.74 | |
endo/exo-crine:10.60 | |
|
| CAGE |
|
brain:6.00 | |
blood:5.88 | |
connective:5.81 | |
reproductive:5.86 | |
muscular:5.54 | |
alimentary:5.74 | |
liver:4.86 | |
lung:5.94 | |
urinary:5.61 | |
endo/exo-crine:6.05 | |
|
| RNA-seq |
|
brain:4.55 | |
blood:6.13 | |
connective:4.96 | |
reproductive:5.93 | |
muscular:4.74 | |
alimentary:4.71 | |
liver:4.38 | |
lung:5.57 | |
urinary:5.57 | |
endo/exo-crine:5.36 | |
|
| Synonym | thiosulfate sulfurtransferase (rhodanese), TST, RDS, MGC19578, Thiosulfate sulfurtransferase, Rhodanese |
|---|
| Refseq ID | NM_003312 |
|---|
| GeneID | 7263 |
|---|
| Unigene ID | Hs.474783 |
|---|
| Probe set ID | 209605_at |
|---|
|
 |
| EST |
|
brain:16.90 | |
blood:5.20 | |
connective:32.10 | |
reproductive:17.80 | |
muscular:46.80 | |
alimentary:161.00 | |
liver:21.10 | |
lung:24.40 | |
urinary:86.40 | |
endo/exo-crine:3.90 | |
|
| GeneChip |
|
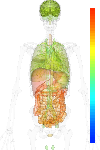
brain:6.94 | |
blood:6.55 | |
connective:7.94 | |
reproductive:7.25 | |
muscular:7.51 | |
alimentary:8.61 | |
liver:10.58 | |
lung:7.15 | |
urinary:9.24 | |
endo/exo-crine:7.56 | |
|
| CAGE |
|
brain:2.37 | |
blood:2.07 | |
connective:2.77 | |
reproductive:2.47 | |
muscular:2.63 | |
alimentary:4.12 | |
liver:5.11 | |
lung:2.74 | |
urinary:2.98 | |
endo/exo-crine:2.52 | |
|
| RNA-seq |
|
brain:2.83 | |
blood:2.43 | |
connective:4.07 | |
reproductive:3.26 | |
muscular:3.44 | |
alimentary:3.62 | |
liver:6.99 | |
lung:2.65 | |
urinary:4.67 | |
endo/exo-crine:3.56 | |
|
| Synonym | HSPC151, HSPC051, HSPC119, UCRC, ubiquinol-cytochrome c reductase complex (7.2 kD) |
|---|
| Refseq ID | NM_001003684 |
|---|
| GeneID | 29796 |
|---|
| Unigene ID | Hs.284292 |
|---|
| Probe set ID | 218190_s_at |
|---|
|
 |
| EST |
|
brain:28.70 | |
blood:70.40 | |
connective:77.10 | |
reproductive:28.70 | |
muscular:312.30 | |
alimentary:30.20 | |
liver:190.00 | |
lung:39.10 | |
urinary:19.20 | |
endo/exo-crine:46.60 | |
|
| GeneChip |
|
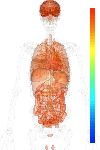
brain:9.80 | |
blood:9.27 | |
connective:9.51 | |
reproductive:9.58 | |
muscular:10.49 | |
alimentary:9.73 | |
liver:9.53 | |
lung:9.23 | |
urinary:10.11 | |
endo/exo-crine:9.44 | |
|
| CAGE |
|
brain:4.90 | |
blood:4.41 | |
connective:4.13 | |
reproductive:4.14 | |
muscular:5.45 | |
alimentary:4.77 | |
liver:5.05 | |
lung:4.54 | |
urinary:4.59 | |
endo/exo-crine:4.50 | |
|
| RNA-seq |
|
brain:2.60 | |
blood:2.96 | |
connective:2.16 | |
reproductive:2.47 | |
muscular:2.87 | |
alimentary:2.67 | |
liver:1.83 | |
lung:2.43 | |
urinary:2.78 | |
endo/exo-crine:2.41 | |
|
| Synonym | HSPC151, HSPC051, HSPC119, UCRC, ubiquinol-cytochrome c reductase complex (7.2 kD) |
|---|
| Refseq ID | NM_013387 |
|---|
| GeneID | 29796 |
|---|
| Unigene ID | Hs.284292 |
|---|
| Probe set ID | 218190_s_at |
|---|
|
 |
| EST |
|
brain:28.70 | |
blood:70.40 | |
connective:77.10 | |
reproductive:28.70 | |
muscular:312.30 | |
alimentary:30.20 | |
liver:190.00 | |
lung:39.10 | |
urinary:19.20 | |
endo/exo-crine:46.60 | |
|
| GeneChip |
|
brain:9.80 | |
blood:9.27 | |
connective:9.51 | |
reproductive:9.58 | |
muscular:10.49 | |
alimentary:9.73 | |
liver:9.53 | |
lung:9.23 | |
urinary:10.11 | |
endo/exo-crine:9.44 | |
|
| CAGE |
|
brain:4.90 | |
blood:4.41 | |
connective:4.13 | |
reproductive:4.14 | |
muscular:5.45 | |
alimentary:4.77 | |
liver:5.05 | |
lung:4.54 | |
urinary:4.59 | |
endo/exo-crine:4.50 | |
|
| RNA-seq |
|
brain:5.79 | |
blood:5.18 | |
connective:4.28 | |
reproductive:5.39 | |
muscular:6.45 | |
alimentary:5.24 | |
liver:5.84 | |
lung:5.17 | |
urinary:6.32 | |
endo/exo-crine:5.24 | |
|
| Synonym | Glycosylation-inhibiting factor, GLIF, MMIF |
|---|
| Refseq ID | NM_002415 |
|---|
| GeneID | 4282 |
|---|
| Unigene ID | Hs.407995 |
|---|
| Probe set ID | 217871_s_at |
|---|
|
 |
| EST |
|
brain:127.50 | |
blood:164.30 | |
connective:128.50 | |
reproductive:162.80 | |
muscular:210.80 | |
alimentary:60.40 | |
liver:52.80 | |
lung:141.70 | |
urinary:172.80 | |
endo/exo-crine:124.30 | |
|
| GeneChip |
|
brain:9.64 | |
blood:9.27 | |
connective:9.22 | |
reproductive:9.91 | |
muscular:8.28 | |
alimentary:9.52 | |
liver:9.60 | |
lung:9.02 | |
urinary:10.42 | |
endo/exo-crine:9.82 | |
|
| CAGE |
|
brain:5.94 | |
blood:5.97 | |
connective:4.98 | |
reproductive:5.54 | |
muscular:6.05 | |
alimentary:5.74 | |
liver:6.46 | |
lung:6.12 | |
urinary:5.78 | |
endo/exo-crine:6.19 | |
|
| RNA-seq |
|
brain:6.32 | |
blood:7.38 | |
connective:7.10 | |
reproductive:7.63 | |
muscular:5.36 | |
alimentary:7.39 | |
liver:6.85 | |
lung:7.33 | |
urinary:7.93 | |
endo/exo-crine:7.43 | |
|
| Synonym | catechol-O-methyltransferase, Catechol O-methyltransferase, COMT |
|---|
| Refseq ID | NM_000754 |
|---|
| GeneID | 1312 |
|---|
| Unigene ID | Hs.370408 |
|---|
| Probe set ID | 208818_s_at |
|---|
|
 |
| EST |
|
brain:54.00 | |
blood:18.30 | |
connective:51.40 | |
reproductive:61.60 | |
muscular:54.70 | |
alimentary:50.30 | |
liver:- | |
lung:9.80 | |
urinary:124.80 | |
endo/exo-crine:77.70 | |
|
| GeneChip |
|
brain:8.44 | |
blood:8.29 | |
connective:8.57 | |
reproductive:8.60 | |
muscular:8.50 | |
alimentary:8.94 | |
liver:10.32 | |
lung:9.42 | |
urinary:9.12 | |
endo/exo-crine:8.75 | |
|
| CAGE |
|
brain:4.07 | |
blood:3.94 | |
connective:3.75 | |
reproductive:4.28 | |
muscular:3.70 | |
alimentary:4.51 | |
liver:5.83 | |
lung:4.54 | |
urinary:4.09 | |
endo/exo-crine:4.02 | |
|
| RNA-seq |
|
brain:2.40 | |
blood:3.76 | |
connective:3.36 | |
reproductive:3.51 | |
muscular:3.01 | |
alimentary:3.57 | |
liver:2.74 | |
lung:3.90 | |
urinary:3.11 | |
endo/exo-crine:3.45 | |
|
| Synonym | catechol-O-methyltransferase, Catechol O-methyltransferase, COMT |
|---|
| Refseq ID | NM_001135161 |
|---|
| GeneID | 1312 |
|---|
| Unigene ID | Hs.370408 |
|---|
| Probe set ID | 208818_s_at |
|---|
|
 |
| EST |
|
brain:54.00 | |
blood:18.30 | |
connective:51.40 | |
reproductive:61.60 | |
muscular:54.70 | |
alimentary:50.30 | |
liver:- | |
lung:9.80 | |
urinary:124.80 | |
endo/exo-crine:77.70 | |
|
| GeneChip |
|
brain:8.44 | |
blood:8.29 | |
connective:8.57 | |
reproductive:8.60 | |
muscular:8.50 | |
alimentary:8.94 | |
liver:10.32 | |
lung:9.42 | |
urinary:9.12 | |
endo/exo-crine:8.75 | |
|
| CAGE |
|
brain:4.07 | |
blood:3.94 | |
connective:3.75 | |
reproductive:4.28 | |
muscular:3.70 | |
alimentary:4.51 | |
liver:5.83 | |
lung:4.54 | |
urinary:4.09 | |
endo/exo-crine:4.02 | |
|
| RNA-seq |
|
brain:1.61 | |
blood:0.00 | |
connective:0.30 | |
reproductive:1.31 | |
muscular:0.00 | |
alimentary:0.00 | |
liver:0.00 | |
lung:0.00 | |
urinary:0.00 | |
endo/exo-crine:0.47 | |
|